Photosymbiosis in freshwater ciliates: elucidating its diversity, functioning and evolution with single-cell sequencing
Symbiosis is best defined as a long-term association of two or more organisms. One of the best-understood symbioses is the partnership between corals and unicellular algae. In this system, the photosynthetic symbiont produces sugars and transfers them into the host, which in return provides shelter to the symbiont. This kind of symbiosis of heterotrophic host and algae is known as photosymbiosis.
Photosymbioses are widespread in marine ecosystems and are highly significant for ecosystem functioning through their important contribution to biomass and primary production. The symbioses enable their participants to overcome the influence of growth-limiting factors by acquiring the scarce substrates from their partners. Among the hosts are animals and microbial eukaryotes (protists), such as radiolarians, foraminiferans, and ciliates. Recently, it has also been shown that symbiotic systems can be much more complex than just host-symbiont pairs – additional microbial players can participate in the functioning and stabilization of symbiosis.
Due to the evolutionary significance of photosymbiosis and its ecological role in aquatic ecosystems, there is a growing number of studies on photosymbiotic systems in marine environments. However, the diversity, functioning and evolution of photosymbiosis in fresh waters are still poorly described, with the bulk of current research focused on a model ciliate Paramecium bursaria and its green algal photosymbiont Chlorella. To improve our understanding of freshwater symbioses, we will focus on different species of photosymbiotic ciliates.

PROJECT OVERVIEW
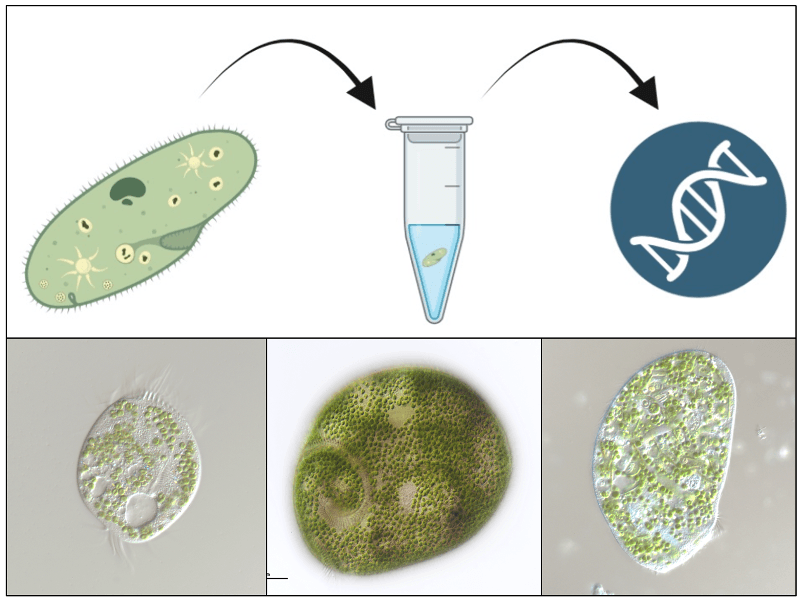
We will use short DNA fragment (molecular barcode) sequencing methods to identify the diversity of photosymbiotic ciliates in selected freshwater environments. We will also collect environmental data, such as nutrient availability, water temperature, dissolved oxygen, and light accessibility, to learn about the environmental preferences of various ciliates of interest. The characterization of the host, symbiont and other possible players will be achieved by sequencing genomes from single cells isolated directly from the environment. Genome analysis will allow for predicting biological interactions between partners, such as the nutrient exchange. We will employ single-cell transcriptome sequencing of hosts and symbionts to analyse their gene expression patterns, which will facilitate the identification of potential mechanisms for controlling host-symbiont interactions.
Carrying out the field experiments will make it possible to evaluate the environmental factors’ influence on the abundance and diversity of freshwater photosymbiotic ciliates. The molecular characterization of the host, symbiont, and other players will help understand the molecular basis of symbiotic systems’ functioning and stability. Results of this project will contribute to our understanding of photosymbiosis and, more broadly, mixotrophy (i.e., the ability to use both heterotrophic and phototrophic modes of nutrition). That is particularly growing in importance due to the role of mixotrophs in aquatic ecosystems under changing environmental conditions. Characterization of the investigated symbiotic systems on a molecular level will also contribute to our understanding of the impact of symbiosis on the evolution of eukaryotes.
SAMPLING
Our main sampling spot are three ponds in Zielonka town near Warsaw which we sampled every month for a year to check how the season affects photosymbioses in ciliates and collect cells for single-cell genomics and transcriptomics. During each sampling we collect water for filtering and chemical analysis plus sediments and concentrated plankton for single-cell picking. We also measure temperature and oxygen level using a multiparametric probe.
Additionally, to cover higher diversity of photosymbiotic ciliates we collect samples from different ponds in Warsaw and in Masurian region.

SINGLE-CELL PICKING
In the lab we inspect collected samples under a light microscope and look for ciliates harboring green algae. We manually pick single cells to perform whole genome amplification and sequence their genomes and transcriptomes.
CONFERENCES
• Chwalińska M., Karnkowska A. Photosymbiosis in freshwater ciliates: elucidating its diversity, functioning and evolution with single-cell sequencing. de.NBI Spring School 2024 – Bioinformatics for Microbial Omics. Bielefeld, Germany, March 18–22, 2024. Poster and flash talk
• Chwalińska M., Karnkowska A. Uncover the nature of ciliate-green algae symbiosis using single-cell genomics. EMBO Early Career Lecture Course “Evolutionary and Comparative Genomics”. Nafplion, Greece, November 3–6, 2024. Poster
• Chwalińska M., Tymoshenko D., Archibald J., Karnkowska A. A single-cell genomic workflow for characterizing unculturable and diverse ciliate–green alga photosymbioses. EMBO | EMBL Symposium: The cellular mechanics of symbiosis: sensing friend from foe. Heidelberg, Germany, March 17-20, 2026. Poster and flash talk
• Chwalińska M., Méndez-Sánchez D., Tymoshenko D., Archibald J., Karnkowska A. Freshwater photosymbiotic ciliates – complex systems reconstructed with single-cell genomics. Protistology Open 2026. Prague, Czech Republic, April 19-23, 2026. Presentation
INTERNSHIPS
PhD Student Małgorzata Chwalińska did a three-month long internship in the research group of Prof. John Archibald in the Institute for Comparative Genomics, Dalhousie University, Halifax, Nova Scotia, Canada (August-November 2025). The internship was sponsored by the Polish National Agency for Academic Exchange NAWA.